How AI can help understand sarcomere structures in hiPSC-CMs?
This year the International Conference for Machine Learning celebrated its 40th anniversary in Honolulu, Hawaii. For those who are busy, here are my 3 key takeaways:
– Healthcare remains one of the dominant domains of application for machine learning;
– Multimodal model building is coming into its own and LLM is supercharging our ability to do this at scale;
– Harmonized data atlases are essential to power AI for biology.
For those with a bit more time, I’ve unpacked each one in more detail below:
1. Healthcare is one of the dominant domains of application for machine learning
ICML is a generalist machine learning conference but this year there was a definite focus on healthcare. Three out of the four keynote talks as well as an unprecedented number of workshops were healthcare-related, exploring topics such as multimodal data for healthcare and computational biology. The most impactful speaker in the space was Prof. Jennifer A. Doudna from the Innovative Genomics Institute, Howard Hughes Medical Institute and University of California Berkeley & UCSF. Her talk highlighted the profound impacts machine learning will have on biological research over the next decade. Drawing from her own experience as a Nobel prize-winning scientist, she explored the intersection of machine learning and CRISPR genome editing and provided exciting examples of the opportunities for AI to help us better understand biology. Doudna explained how machine learning may in the near future accelerate and perhaps fundamentally alter our use of CRISPR in both humans and microbes.
 Image of Dr Jennifer Doudna, recipient of the 2020 Nobel Prize in chemistry with the Cas9 protein (white) which is shown in surface format and the guide RNA (orange) and target DNA (blue). Source: https://3dmoleculardesigns.com/custom_models/crispr/.
Image of Dr Jennifer Doudna, recipient of the 2020 Nobel Prize in chemistry with the Cas9 protein (white) which is shown in surface format and the guide RNA (orange) and target DNA (blue). Source: https://3dmoleculardesigns.com/custom_models/crispr/.
2. Multimodal model building is coming into its own and LLM is supercharging our ability to do this at scale
Multimodalism was a major topic at ICML. A key limitation to building robust multimodal models in healthcare remains availability of high quality, labelled data that is harmonized and AI-ready. A potential remedy proposed by Deldari et al is self-supervised learning like CroSSL (Cross-modal SSL) which makes intermediate embeddings from modality-specific encoders and aggregates them into a global embedding using a cross-modal aggregator.
Large language models (LLMs) continue to show an impressive ability to solve tasks in healthcare applications. But in order for LLMs to effectively solve personalized health tasks, LLMs need the ability to ingest a diversity of data modalities that are relevant to an individual’s health status. A paper by Cosentino et al took steps towards creating multimodal LLMs for health that are grounded in individual-specific data by developing a framework (HeLM: Health Large Language Model for Multimodal Understanding) that enables LLMs to use high-dimensional clinical modalities to estimate underlying disease risk. HeLM encodes complex data modalities by learning an encoder that maps them into the LLM’s token embedding space and, for simple modalities like tabular data, by serializing the data into text. Using data from the UK Biobank, the authors showed that HeLM can effectively use demographic and clinical features in addition to high dimensional time-series data to estimate disease risk. For example, HeLM achieves an AUROC of 0.75 for asthma prediction when combining tabular and spirogram data modalities compared with 0.49 when only using tabular data.
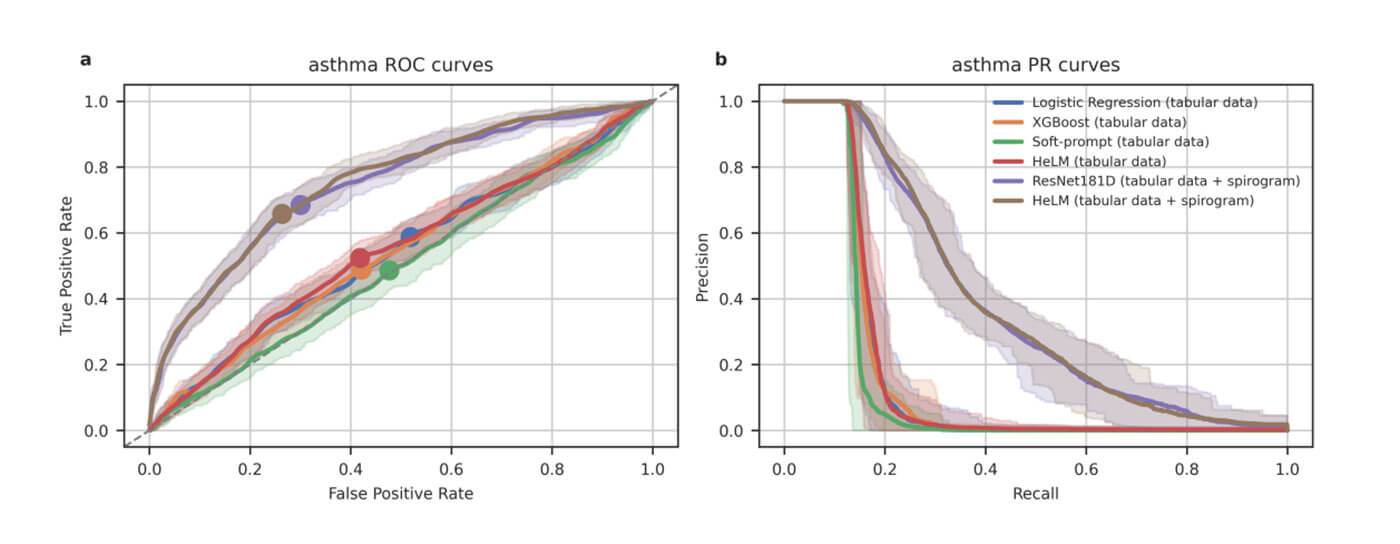 Inclusion of additional input modalities improves HeLM asthma detection. Source: Belyaeva, A., Cosentino, J., Hormozdiari, F., McLean, C.Y. and Furlotte, N.A., 2023. Multimodal LLMs for health grounded in individual-specific data. arXiv preprint arXiv:2307.09018.
Inclusion of additional input modalities improves HeLM asthma detection. Source: Belyaeva, A., Cosentino, J., Hormozdiari, F., McLean, C.Y. and Furlotte, N.A., 2023. Multimodal LLMs for health grounded in individual-specific data. arXiv preprint arXiv:2307.09018.
3. Data atlases are essential to power machine learning
There was consensus at ICML that building biological data atlases from healthy and diseased tissue is the first and very valuable step in accelerating our ability to understand complex biology through AI. Deepmind’s AlphaFold is a perfect example of this: AlphaFold required a vast data atlas of experimentally-determined protein structures from the Protein Data Bank which had been painstakingly prepared by thousands of researchers over many years.
Many researchers at ICML believe that spatial omics and single-cell technologies will be the next frontier in which a large atlas will unlock medical breakthroughs. Research shared at ICML by Fabian Theis integrated many single-cell datasets to address the limitations of individual studies to capture the variability present in a given population. He presented the integrated Human Lung Cell Atlas (HLCA), which combines 49 datasets of the human respiratory system into a single atlas spanning over 2.4 million cells from 486 individuals. Leveraging the number and diversity of individuals in the HLCA, the researchers identified gene modules (i.e. sets of genes with similar expression profiles) that are associated with demographic covariates such as age, sex and body mass index, as well as gene modules changing expression along the proximal-to-distal axis of the bronchial tree. Mapping new data to the HLCA enabled rapid data annotation and interpretation. Using the HLCA as a reference for the study of disease, they identified shared cell states across multiple lung diseases, including SPP1+ profibrotic monocyte-derived macrophages in COVID-19, pulmonary fibrosis and lung carcinoma.
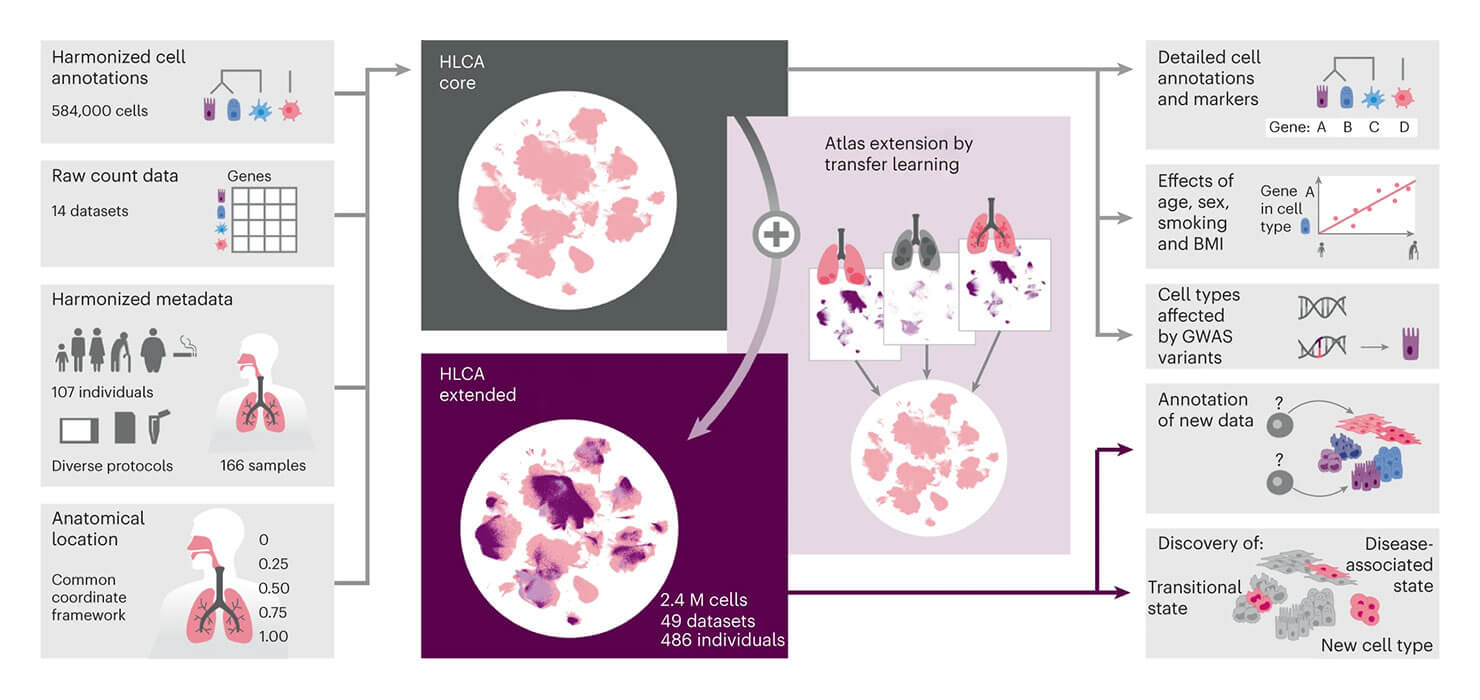 Source: Sikkema, L., Ramírez-Suástegui, C., Strobl, D.C. et al. An integrated cell atlas of the lung in health and disease. Nat Med 29, 1563–1577 (2023). https://doi.org/10.1038/s41591-023-02327-2.
Source: Sikkema, L., Ramírez-Suástegui, C., Strobl, D.C. et al. An integrated cell atlas of the lung in health and disease. Nat Med 29, 1563–1577 (2023). https://doi.org/10.1038/s41591-023-02327-2.
This is excellent news for Owkin’s recently launched MOSAIC initiative. MOSAIC is Owkin’s landmark $50 million project to revolutionize cancer research through the use of spatial omics, a set of cutting-edge technologies that offer unprecedented information on the structure of tumors. By generating and analyzing unprecedented amounts of spatial omics data in combination with multimodal patient data and artificial intelligence, MOSAIC aims to unlock the next wave of treatments for some of the most difficult-to-treat cancers. Collaborating with Gustave Roussy, the University of Pittsburgh through the UPMC Hillman Cancer Center, Lausanne University Hospital, Uniklinikum Erlangen/Friedrich-Alexander- Universität Erlangen-Nürnberg, Charité – Universitätsmedizin Berlin, and providers of analytical instrumentation and assays for spatial biology. MOSAIC will use 7,000 tumor samples from patients, making it over 100 times larger than any existing spatial omics datasets. Owkin and the MOSAIC partners will mine this resource for immune-oncology disease subtypes in pursuit of biomarkers and novel therapies.
This is an article from Owkin – an AI Biotech that uses AI to unlock complex biology to find the right treatment for every patient. Re-posted by VISHC News. The original post can be found here.